Histone mRNA-cleaving machine, crucial for healthy cell division, yields up its secrets after almost half a century of research. Zbigniew Dominski, PhD, and William Marzluff, PhD, led this effort at the UNC School of Medicine, along with key collaborators at Columbia University and Rockefeller University.

Histone mRNA-cleaving machine, crucial for healthy cell division, yields up its secrets after almost half a century of research. Zbigniew Dominski, PhD, and William Marzluff, PhD, led this effort at the UNC School of Medicine, along with key collaborators at Columbia University and Rockefeller University.


CHAPEL HILL, NC – February 6, 2020 [under embargo until 2 pm EST] – Scientists from the UNC School of Medicine, Columbia University, and Rockefeller University have revealed the inner workings of one of the most fundamental and important molecular machines in cells.
The researchers, in a study published in Science, used biochemical experiments and cryo-electron microscopy (cryo-EM) to determine the atomic structure of a complex assembly of molecules known as the histone mRNA three-prime (3’) end-processing machine. This machine plays a fundamental role in proper activity and duplication of the cell genome and when defective, it may lead to human diseases, including cancers.
Histone proteins are found in all plants and animals, and they form a “beads-on-a-string” arrangement where the DNA in chromosomes is wrapped around the beads of histones. Histones ensure the efficient packaging of DNA and help regulate which genes are turned “on” and which are kept “off,” processes needed for all cells to function properly.
The histone mRNA 3’ end-processing machine is responsible for cutting – at precisely the right place – the mRNA transcript that is copied out from a histone gene and encodes the corresponding histone protein. The machine performs an essential role in cells’ production of histone proteins, which occurs at high levels whenever a cell divides and must replicate its DNA. The structure shows how the machine is activated only after it binds the histone mRNA, preventing cleavage of other RNAs.
“This structure provides the first atomic insights into a critical process in cells, and beautifully explains the large body of current knowledge on this machinery,” said senior author Liang Tong, PhD, William R. Kenan, Jr. Professor of the Biological Sciences Department at Columbia University. “The structure has been long awaited by scientists in the field, and the elegant amphora shape of the machinery is an unexpected bonus. The structure also provides valuable insights into other RNA 3’-end processing machineries, because they share key components with the histone machinery.”
Tong’s recent studies showed the canonical machinery in an inactive form. So, now, scientists have a glimpse of how the machinery is activated. “This structure is another illustration of the remarkable power of the new cryo-EM technique,” Tong added.
The solution of the structure of this complex, a landmark achievement of molecular biology, is the culmination of nearly 40 years of research by a number of laboratories and molecular biologists.
“I started on the problem of histone mRNA and how it’s regulated when I first started my laboratory as an assistant professor at Florida State in 1974,” said study co-author William Marzluff, PhD, Kenan Distinguished Professor of Biochemistry and Biophysics at the UNC School of Medicine and the Integrative Program in Biological and Genome Sciences at UNC-Chapel Hill. “And this is certainly the most important contribution we have made in this field of inquiry so far.”
“For a long time we have been studying different pieces of this molecular machine, but now for the first time we know how all the pieces fit together and work together,” said Zbigniew Dominski, PhD, professor in the Department of Biochemistry & Biophysics at the UNC School of Medicine, who led the discovery of many of the components of the machine. “It’s as if someone opened up the hood of an old car so that at last you could see how the whole engine looks and works, suddenly learning about unexpected mechanical and functional details.”
Dominski was a co-corresponding author of the study with Thomas Walz, PhD, Professor and Head of Laboratory of Molecular Electron Microscopy at Rockefeller University.
 A special tail
A special tail
Every protein is produced in a process that starts with a gene. Special enzymes copy out, or transcribe, the information in the gene in the form of ribonucleic acid (RNA), a close molecular cousin of DNA in the cell nucleus. A special molecular machine called a 3’ end processing machine must then cut that strand of RNA at the correct place to process it into a molecule called a messenger RNA (mRNA), which migrates into the main part of the cell and is translated there into the final protein.
The mRNAs for virtually all proteins are processed by one type of 3’ end-processing machine, which cuts them at the correct place and adds a special tail to them. Histone transcripts in animal cells which encode histone proteins needed for cell division are processed by a different machine, which cuts them but adds no tail. And this is the machine that we are now very familiar with due to this breakthrough structural study.
“No one really knows why histone mRNAs are different from other mRNAs; it’s what we call a theological question,” Marzluff joked.
The canonical and histone RNA 3’ end-processing machines are each composed of more than a dozen individual proteins and RNA molecules. Some of these elements are found in both machines, suggesting a common evolutionary origin. Since the histone 3’ processing machine contains the same three core proteins as the canonical machine, including the protein that actually cleaves the RNA, the process of activation of the two machines is likely similar, although the way the two machines recognize their RNA targets is distinct.
Tong, Dominski, Marzluff and their colleagues succeeded in assembling a working version of the histone RNA 3’ end-processing machine from its 13 protein and 2 RNA components, essentially in a test-tube. The machine was then imaged using cryo-electron microscopes at the New York Structural Biology Center (NYSBC), and subsequent data processing ultimately led to a structure at near atomic resolution. The team was also able to mutate key components to verify their individual functions.
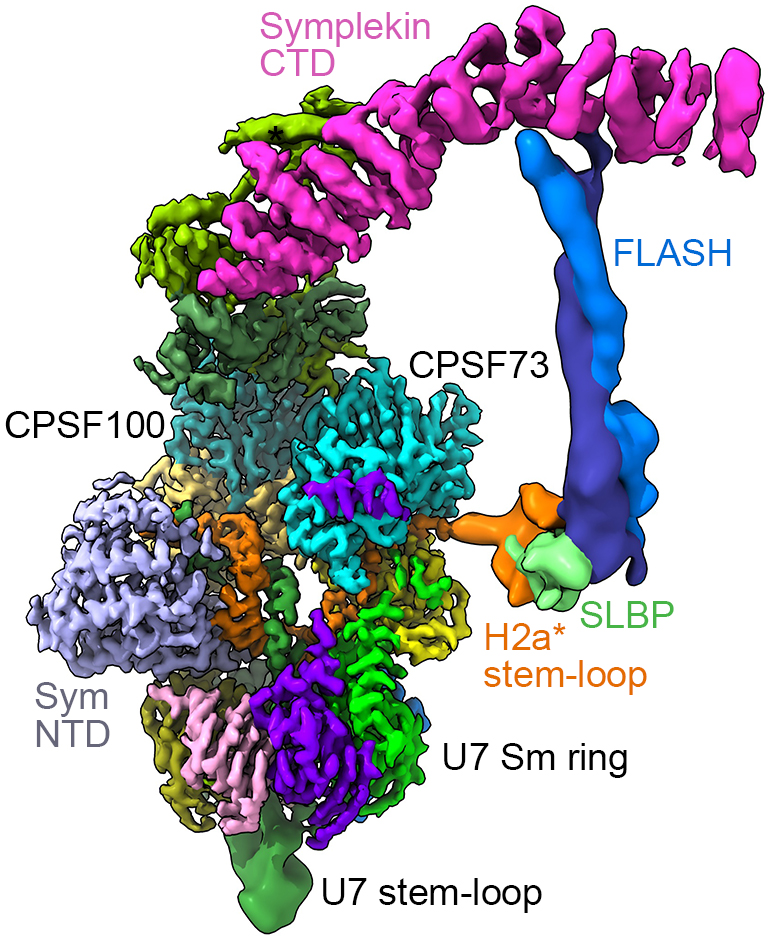
The amphora
The structure of the machine turned out to resemble an amphora with one long handle. The cryo-EM analysis also revealed how the machine recognizes histone RNA and cuts it at precisely the right place.
“It detects two elements of the RNA strand, and only when they are present does the cutting device in this machine expose its blades, so to speak,” Dominski said. “There is no randomness, no accident; it cleaves only what it is supposed to cleave, and the study reveals this beautifully.”
Dominski has been investigating this histone RNA-processing machinery since the mid-1990s and with Marzluff, Tong, and others has been responsible for key discoveries about individual components. “This is more or less the end of the road, as far as understanding how this machine works,” he said. “We’ve resolved that pretty clearly with this study, and it’s a good feeling.”
Other authors were first authors Yadong Sun of Columbia University and Yixiao Zhang of Rockefeller University, Wei Shen Aik of Columbia, and Xiao-Cui Yang of UNC-Chapel Hill. Marzluff is a member of the UNC Lineberger Comprehensive Cancer Center.
The National Institutes of Health funded this research. Read more about the importance of cryo-EM here.
Media contacts:
Mark Derewicz, UNC School of Medicine, 984-974-1915
Carla Cantor, Columbia University, 212-854-5276